A model of the influenza genome architecture—untouched since the 1970s—isn’t so perfect after all. Scientists are ready to give it an overhaul.
New research reveals loopholes in the way the virus packages its genetic material. When one strain of flu co-mingles with another strain inside a cell, the loopholes allow the viruses to swap genetic material and give rise to new strains of flu.
Knowing about the loopholes and how they interact with each other could offer the opportunity to better predict pandemics and find new ways to disrupt the flu virus.
“Although influenza has plagued mankind for hundreds of years and poses a substantial public health threat every winter, we know surprisingly little about flu pandemics,” says Seema S. Lakdawala, assistant professor of microbiology & molecular genetics at the University of Pittsburgh and senior author of the study in Nucleic Acids Research.
“Our discovery may give insight into how the flu virus continually evolves, opening the door to better vaccines and antivirals.”
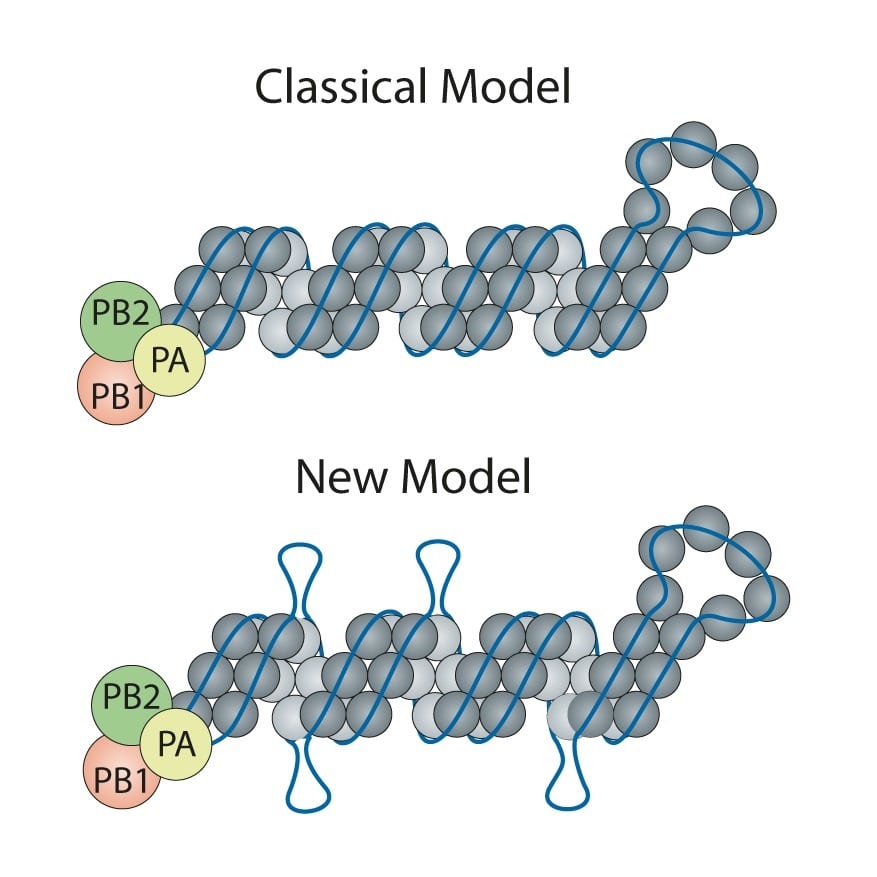
Influenza is a type of virus that uses single-stranded ribonucleic acid (RNA) to replicate, instead of double-stranded DNA. Influenza viruses are made up of eight RNA segments bound by a protective nucleoprotein. All eight RNA segments must come together inside a virus particle to be fully infectious.
The classic model of the flu virus has these proteins coating the RNA like beads evenly spaced along a string. However, limitations of techniques used in the 1970s when the model was developed meant that unique features—like exposed RNA loops—were lost. Consequently, the universal depiction of influenza in textbooks is of a uniform random binding of proteins along the entire length of each RNA segment.
Researchers were curious if there might be any areas along the influenza RNA strand that are more “open” and, therefore, more able to associate with other RNA segments in order to arrive at a package of all eight segments.
They used a process called “high-throughput sequencing of RNA by crosslinking immunoprecipitation” (HITS-CLIP) on two strains of influenza A, including the 2009 H1N1 pandemic strain, to get a better understanding for where the proteins bind to the RNA and to see if there were any areas of “naked” RNA.
Outbreaks hinge on sick people caring about others
“Honestly, we didn’t expect to find any since we had all learned the ‘beads on a string’ depiction of viral RNA,” Lakdawala says. “But, amazingly, there are several stretches where the RNA was not bound by the nucleoprotein. This discovery opens up a whole new area of research.”
Contrary to the classic model, Lakdawala and coauthor author Nara Lee, assistant professor of microbiology & molecular genetics found there are areas of RNA rich with protein coating and others that are exposed and presumably ripe for binding to other viral RNA during re-assortment, or the swapping of genomic material between flu viruses.
Painless patch works just as well as flu shot
The team is pursuing several potential research opportunities, including predicting the ways different influenza viruses could share genetic material to make new viruses. Knowing this could point scientists to the re-assortments most likely to spark a flu pandemic and give public health agencies a leg-up on creating targeted vaccines. There also could be ways to exploit the exposed RNA to make the virus less transmissible and deadly.
“It’s really exciting to suddenly have all these research possibilities open up based on this one discovery,” Lakdawala says. “The reason no one’s uncovered this yet is because we all took for granted that 50-year-old research on the genome architecture, which looked really nice and had an easy explanation, was the full story. It shows that if we don’t constantly resample and question scientific dogma, we could miss a big opportunity.”
The University of Pittsburgh and National Institutes of Health supported the work.
Source: University of Pittsburgh