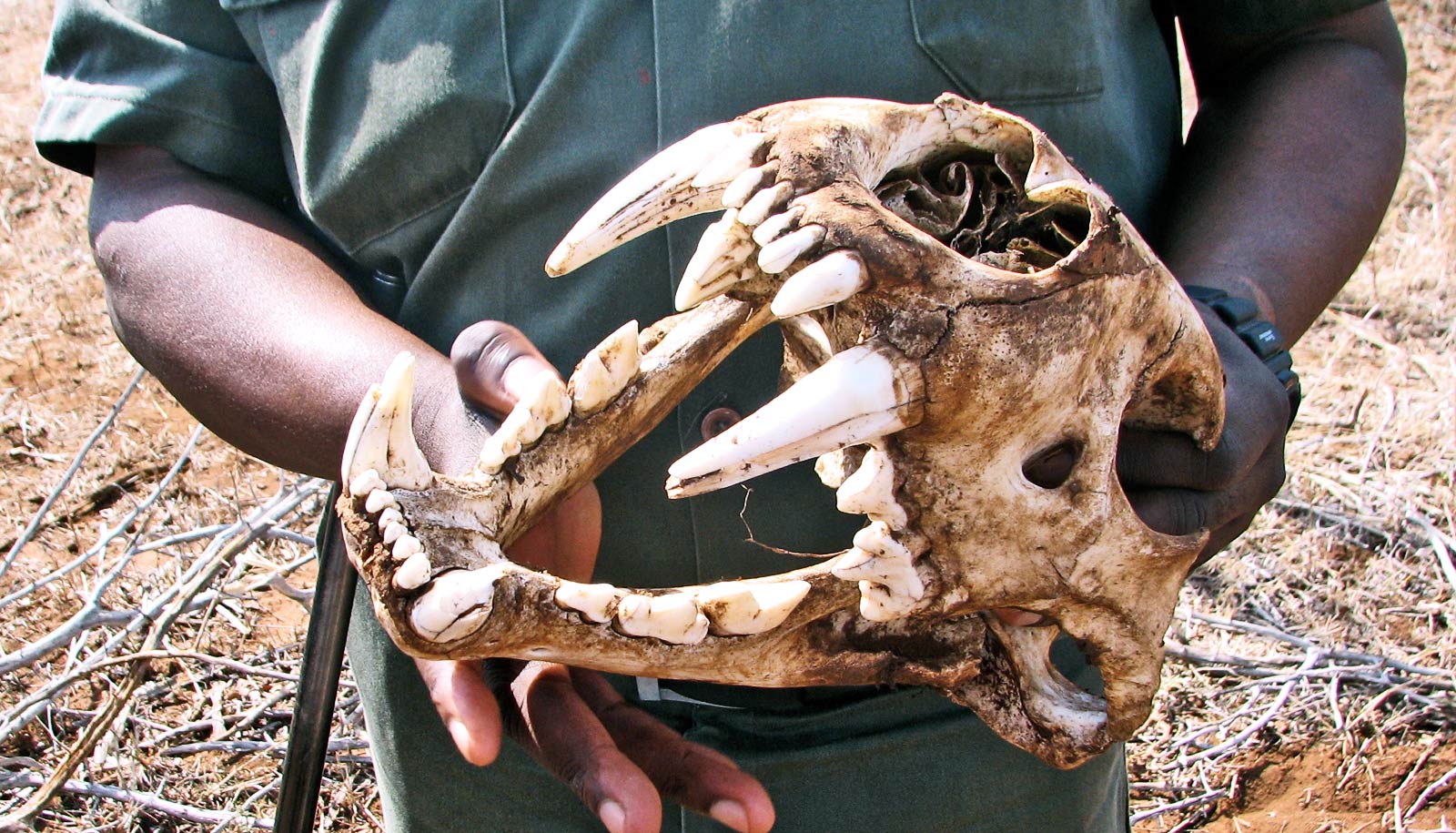
Scientists have come up with a new way to collect DNA from endangered species—extract it from degraded and left-behind materials, including feces, saliva, and even food products.
The near impossibility of collecting DNA samples from rare and elusive animals has hobbled wildlife detectives aiming to protect these endangered species. The researchers say their proof of concept, which appears in Methods in Ecology and Evolution, could revolutionize conservation approaches and policies worldwide.
“It’s CSI meets conservation biology,” says coauthor Dmitri Petrov, professor in the School of Humanities and Sciences at Stanford University.
Looming extinctions
The specter of extinction hangs over more than a quarter of all animal species, according to the best estimate of the International Union for Conservation of Nature, which maintains a list of threatened and extinct species. Conservationists have documented extreme declines in animal populations in every region of Earth.
“Conservation needs answers fast, and our research was not providing them fast enough.”
Helping species recover often depends on collecting DNA samples, which can reveal valuable information about details including inbreeding, population history, natural selection, and large-scale threats such as habitat destruction and illegal wildlife trade.
However, current approaches tend to require relatively large amounts of DNA or expensive and often inefficient strategies for extracting the material. Getting meaningful information rapidly from lower-concentration, often degraded and contaminated DNA samples requires expensive and specialized equipment.
Tigers and conchs
A solution may lie in an ongoing collaboration between Stanford’s Program for Conservation Genomics, including the labs of Petrov and coauthors Elizabeth Hadly and Stephen Palumbi, and India’s National Centre for Biological Sciences, including the lab of coauthor Uma Ramakrishnan, a molecular ecologist and former Fulbright faculty fellow at Stanford.
“I have been working on tiger conservation genetics for over a decade, but have been frustrated at how slow and unreliable the process of generating genetic data can be,” Ramakrishnan says. “Conservation needs answers fast, and our research was not providing them fast enough.”
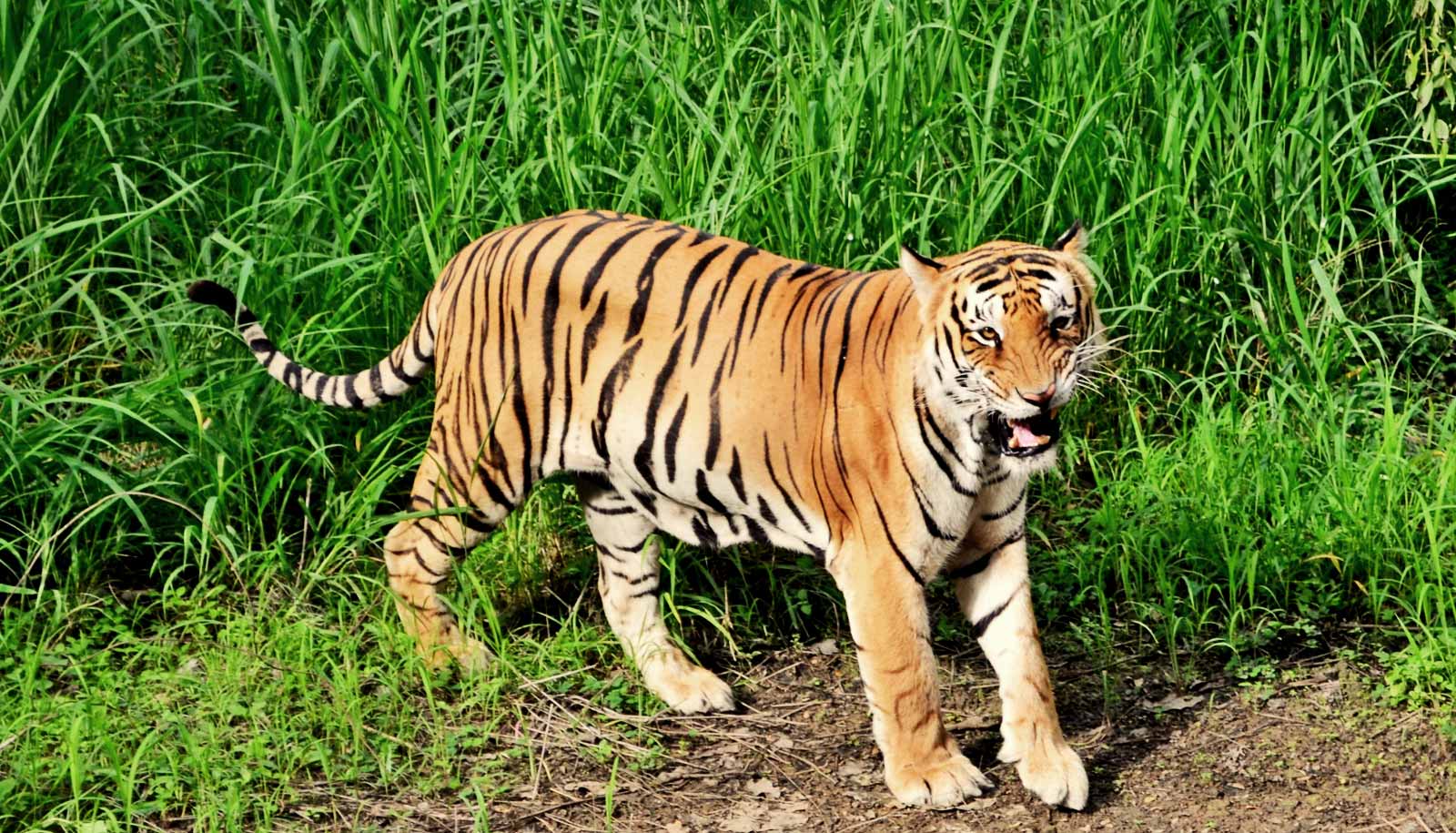
The researchers looked at endangered wild tigers in India and overfished Caribbean queen conchs, examining tiger feces, shed hair, and saliva found on killed prey, as well as fried conch fritters purchased in US restaurants. All of the samples were too impure, mixed, or degraded for conventional genetic analysis.
“Our goal was to find extremely different species that had strong conservation needs, and show how this approach could be used generally,” says Palumbi, professor of marine biology. “The King of the Forest—tigers—and Queen of the Caribbean—conch—were ideal targets.”
Tiny bits of DNA
Together, the team improvised a new approach, using a sequencing method that amplifies and reads small bits of DNA with unique differences in each sample. Doing this simultaneously across many stretches of DNA in the same test tubes allowed the researchers to keep the total amount of DNA needed to a minimum.
Making the procedure specific to tiger and conch DNA allowed for the use of samples contaminated with bacteria or DNA from other species.
The technology proved highly effective at identifying and comparing genetic characteristics. For example, the method worked with an amount of tiger DNA equivalent to about one-one-hundred-thousandth the amount of DNA in a typical blood sample. The method had a higher failure rate in conchs because the researchers did not have whole genomes at their disposal.
The approach’s effectiveness, speed and affordability—implementation could cost as little as $5 per sample—represents a critical advance for wildlife monitoring and forensics, field-ready testing, and the use of science in policy decisions and wildlife trade, the researchers say.
“It is easy to implement and so can be done in labs with access to more or less basic equipment,” says coauthor Meghana Natesh of the National Centre for Biological Sciences and Sastra University in India. “If a standard procedure is followed, the data generated should be easy to share and compare across labs. So monitoring populations across states or even countries should be easier.”
The scientists have made their methods freely available. “We are working to expand the method so that it can identify other species and other characteristics, such as diet and pathogens,” Hadly says.
The Wildlife Conservation Trust, the US Department of State, the Wellcome Trust / DBT India Alliance, the Summit Foundation, and the Smithsonian Institution funded the work, which appears in
Methods in Ecology and Evolution.
Source: Stanford University