New, license-free DNA ladders offer a much cheaper way to estimate the size of DNA fragments.
Researchers developed two plasmids—a circular form of DNA—that DNA scissors known as restriction enzymes can cut to create the DNA ladders. The ladders can be used to estimate the size of DNA fragments between about 50 and 5,000 base pairs in length.
“DNA ladders, also known as DNA molecular weight markers, are among the most commonly used reagents in molecular biology research,” says Song Tan, professor of biochemistry and molecular biology at Penn State. “They are used in any application that requires gel electrophoresis—a technique that separates fragments of DNA by their size.
“We would like to offer these plasmids to the research community as a means to produce high quality DNA molecular weight markers at a low cost.”
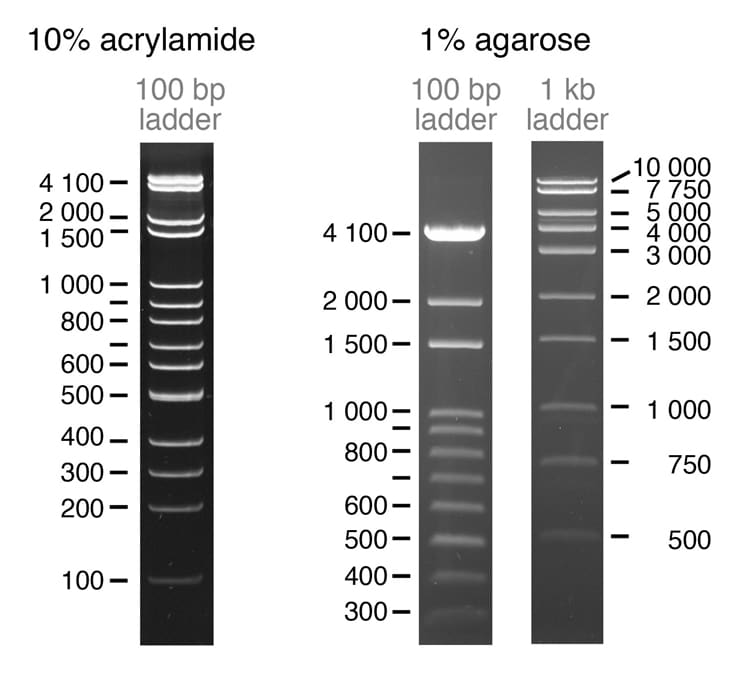
The research team created two plasmids, pPSU1 and pPSU2, that together produce DNA ladders in increments of either 100 or 1,000 base pairs, depending on which restriction enzyme is used. Researchers can easily produce in their own laboratories enough of the two ladders for 1,000 uses for under $10. In contrast, commercially available DNA ladders cost between $250 and $500 for the same amount.
Additionally, unlike many currently available DNA ladders, the 100-base-pair ladders work appropriately on both agarose and polyacrylamide gels, two types commonly used in molecular biology.
“We are also excited about the possibility that the pPSU plasmids may be used around the world to further research and enhance science education in classroom laboratories,” says former undergraduate student Ryan C. Henrici. “This technology produces DNA ladders at less than a penny per use, a fraction of the cost of using commercially available DNA ladders.”
How just 13 DNA snippets could identify you
The pPSU1 and pPSU2 plasmids used to produce the Penn State DNA ladders will be available without licensing restrictions to nonprofit academic users through the Addgene and DNASU plasmid repositories.
The US National Institutes of Health – National Institute of General Medical and the Penn State Eberly College of Science supported the work. A paper describing the research appears in Scientific Reports.
Source: Sam Sholtis for Penn State



